Generation of a P-iKO hESC line
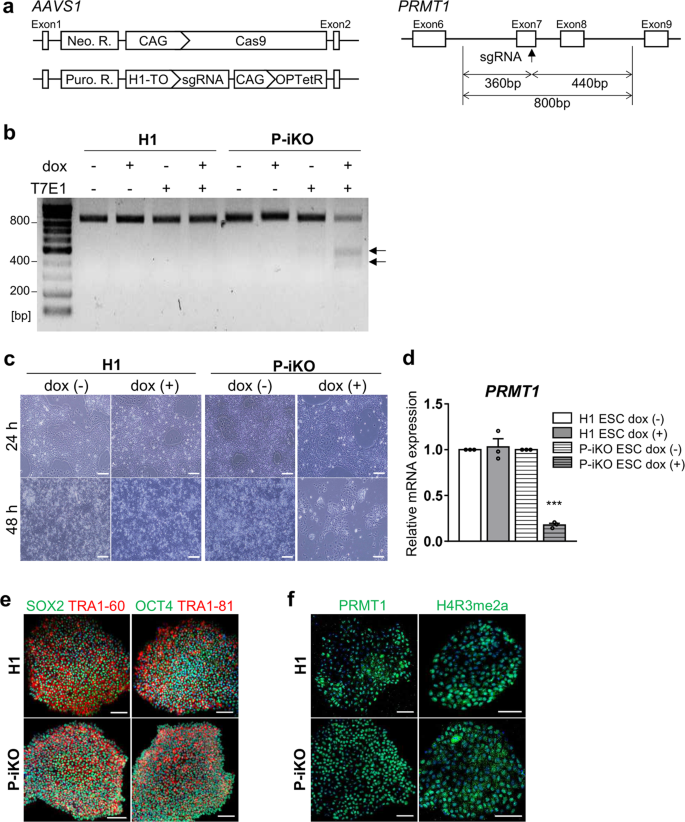
To understand the role of arginine methylation in pancreatic endocrine cell development, we generated a PRMT1-inducible knockout (P-iKO) hESC line (Supplementary Fig. 1). One AAVS1 allele was targeted for constitutive Cas9 expression, and the other was targeted for tetracycline-dependent (TetR) sgRNA transcription (Fig. 1a, left panel). We designed the sgRNA to target human PRMT1 exon 7, which encodes the middle part of the SAM-dependent methyltransferase domain (Fig. 1a, right panel). In normal media, P-iKO cells maintain an intact PRMT1 genomic locus because they produce only the Cas9 protein. The P-iKO cells begin expressing the sgRNA when exposed to doxycycline (dox). Then, the Cas9-sgRNA complex makes indels in the PRMT1 locus of P-iKO cells (Fig. 1b, arrows: 360, 440 bp). In the presence of dox, P-iKO hESCs died within 48 h, whereas H1 hESCs survived (Fig. 1c). As expected, PRMT1 transcript levels were significantly lower in dox-treated P-iKO hESCs than in untreated ones (Fig. 1d). However, similar to H1 hESCs, P-iKO hESCs without dox treatment exhibited normal expression of pluripotency markers, including SOX2, TRA 1-60, OCT4, and TRA 1–81 (Fig. 1e). They also showed normal expression of PRMT1 and methylation of histone 4 (H4R3me2a) in the absence of dox (Fig. 1f). Thus, we generated pluripotent P-iKO hESCs with an inducible P-KO system in vitro.
a AAVS1 locus-targeted alleles for the generation of the P-iKO hESC line (left). Schematic of the PRMT1 exon targeted using the doxycycline-inducible CRISPR system (right). Neo. R., Neomycin resistance gene; Puro. R., puromycin resistance gene; H1-TO, tetracycline-inducible H1 RNA Polymerase III promoter containing a tetO2 sequence; OPTetR, OPTimized Tetracycline-responsive Repressor28. See also Supplementary Fig. 1. b T7E1 analysis of genomic DNA from P-iKO hESCs. A single transgenic cell line after dox treatment was identified by the two genomic DNA fragments indicated by arrows (at 360 and 440 bp). c P-iKO hESC morphology. Most P-iKO hESCs died within 48 h of dox treatment. Scale bars, 200 μm. d Relative PRMT1 mRNA levels in P-iKO hESCs. The relative expression values are presented as the mean ± SEM (n = 3). ***p < 0.001. e Normal expression of pluripotency-associated markers in P-iKO hESCs. Scale bars, 100 μm. f Immunostaining of PRMT1 and H4R3me2a in P-iKO hESCs. Scale bars, 100 μm.
P-iKO hESCs were differentiated into pancreatic endocrine cells in the absence of doxycycline
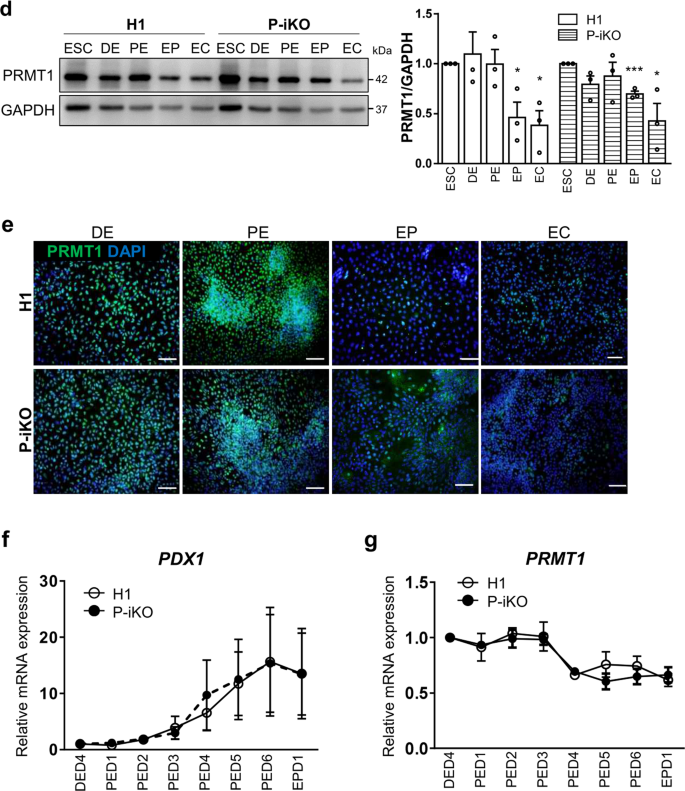
The protocol we used to differentiate human ESCs into pancreatic endocrine cells (ECs) is depicted in Fig. 2a. We found that H1 and P-iKO hESCs both differentiated normally into definitive endoderm (DE), pancreatic endoderm (PE), endocrine progenitor (EP), and ECs according to appearance (Supplementary Fig. 2a). In the absence of dox, we did not observe any difference between H1 and P-iKO hESCs in the transcription of genes associated with each developmental stage (Fig. 2b). We also observed similar expression of the same markers in each developmental stage in an immunostaining in H1 and P-iKO hESCs (Fig. 2c and Supplementary Fig. 2b). In the absence of dox, the expression of PRMT1 was similar in each group, falling significantly from the EP stage (Fig. 2d, e). To determine the timing of dox treatment for P-KO induction, we examined PDX1 expression daily. In both H1 and P-iKO hESCs, PDX1 transcript levels gradually increased from Day 3 of PE stage (PED3) and then decreased upon reaching the EP stage (Fig. 2f). During the same period, the transcription profiles of PRMT1 also remained similar in both groups (Fig. 2g). Thus, after confirming the similarity of H1 and P-iKO hESCs in the absence of dox, for all subsequent experiments, we added dox to the PE medium beginning on PED3 for an additional 3 d to knockout PRMT1 (Supplementary Fig. 2c). After dox treatment, P-iKO cells had normal morphologies at the PE and EP stages but showed relatively flattened shapes at the EC stage compared to those of the non-treated cells (Supplementary Fig. 2d). In addition, dox treatment did not influence the cell number or viability in P-iKO PEs, nor did it influence the cell cycle or the number of EPs (Supplementary Fig. 2e, f). Collectively, these results confirm that pluripotent P-iKO hESCs differentiate into pancreatic ECs.
a Overall protocol for the differentiation of hESCs into pancreatic endocrine cells. The detailed procedures are described in the “Materials and methods” section. D, day. See also Supplementary Fig. 2a. b Transcript expression of developmental genes in P-iKO hESCs in the absence of dox treatment during pancreatic differentiation. The relative expression values are presented as the mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001 (n = 3–4). See also Supplementary Fig. 2b. c Immunostaining of pancreatic developmental markers in P-iKO hESCs undergoing pancreatic differentiation. Scale bars, 100 μm. d Expression of PRMT1 protein in P-iKO hESCs as they progressed through the stages of pancreatic differentiation. The relative expression values are presented as the mean ± SEM. *p < 0.05, ***p < 0.001 (n = 3). e Immunostaining of PRMT1 in P-iKO hESCs as they progressed through the stages of pancreatic lineage differentiation. Scale bars, 100 μm. f Transcriptional profiles of the PDX1 gene as P-iKO hESCs and controls progressed from Day 4 of the DE stage (DED4) to Day 1 of the EP stage (EPD1) of pancreatic lineage differentiation. The data are presented as the mean ± SEM (n = 6). g Transcriptional profiles of the PRMT1 gene in P-iKO hESCs and controls progress from Day 4 of the DE stage (DED4) to Day 1 of the EP stage (EPD1) of pancreatic lineage differentiation. The data are represented as the mean ± SEM (n = 5).
PRMT1-KO impairs the function of NGN3 in pancreatic endocrine development
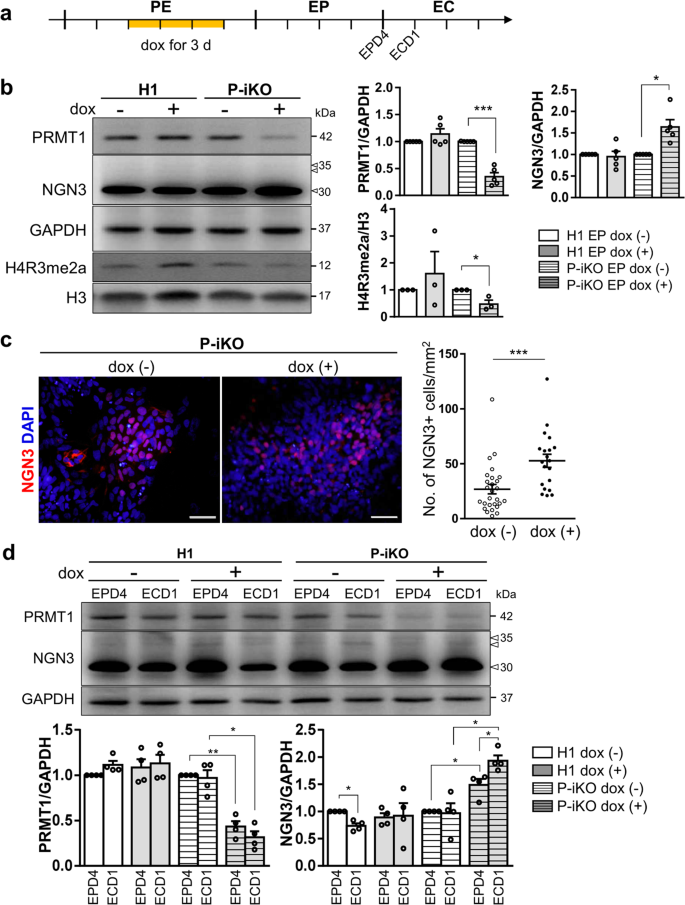
NGN3 is an essential transcription factor for endocrine cell fate decisions and has a very short half-life31. We confirmed the EP stage-specific NGN3 expression in both H1 and P-iKO hESCs during in vitro pancreatic endocrine differentiation, observing robust expression only in the EP stage that fell abruptly by the EC stage (Supplementary Fig. 3a). To determine whether P-KO was also associated with NGN3 accumulation in this study, we treated P-iKO cells with dox in the PE for 3 d and further differentiated them into EPs (Fig. 3a). In the presence of dox, we observed reduced expression of PRMT1 (p < 0.001) but enhanced expression of NGN3 (p < 0.05) in EPs (Fig. 3b). We observed reduced H4R3me2a in P-KO cells compared to untreated cells (Fig. 3b) and more NGN3-positive P-KO cells than untreated P-iKO cells (p < 0.001, Fig. 3c). This increase in NGN3-positive P-KO cells continued into the EC stage (Supplementary Fig. 3b, c). Thus, we demonstrated that P-KO increases the proportion of NGN3-positive cells in hESCs undergoing pancreatic development.
a Schedule for the dox treatment of PEs differentiated from P-iKO hESCs (yellow line). Dox was administered for 3 d to PRMT1-iKO hESCs that had progressed to D3 of the PE stage of pancreatic lineage development. See also Supplementary Fig. 2d–f. b Expression of PRMT1 and NGN3 in P-iKO EPs treated with dox. Relative PRMT1 and NGN3 protein expression levels were normalized to GAPDH levels. The H4R3me2a expression levels were normalized to H3 levels. The data are presented as the mean ± SEM. *p < 0.05, ***p < 0.001 (n = 3–5). See also Supplementary Fig. 3a. c Immunostaining of NGN3 in P-iKO EPs treated with dox (left). NGN3-positive cells were counted using ImageJ (right). P-KO EPs accumulate NGN3. The data are presented as the mean ± SEM. ***p < 0.001. Dox(−), n = 27; dox(+), n = 20. Scale bars, 50 μm. See also Supplementary Figs. 3–5. d Expression of PRMT1 and NGN3 in P-iKO EPs treated with dox. Western blot analysis showed reduced PRMT1 and enhanced NGN3 levels in P-KO EPD4 and ECD1 cells. PRMT1 and NGN3 protein expression levels were normalized to those of GAPDH. Dox(+) and dox(-) indicate treated and untreated cells, respectively. The data are presented as the mean ± SEM. *p < 0.05, **p < 0.01 (n = 4). e Relative mRNA expression of EP-associated genes in P-KO EPs. The transcription of NGN3 target genes was significantly downregulated in P-KO EPs. The data are presented as the mean ± SEM. *p < 0.05, ***p < 0.001 (n = 4). See also Supplementary Fig. 3d–g. f Relative mRNA expression of EC-associated genes in P-KO ECs. Transcripts of NGN3 target genes and EC-associated genes were significantly decreased in P-KO ECs. The data are presented as the mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001 (n = 3). See also Supplementary Fig. 3h, i. g Relative mRNA expression of endocrine hormone genes in P-KO ECs. INS and SST genes were transcriptionally downregulated in P-KO ECs. The data are presented as the mean ± SEM. ***p < 0.001 (n = 4). INS, insulin; PPY, pancreatic polypeptide; SST, somatostatin. h Immunostaining of endocrine hormones in P-KO ECs. GCG-positive cells were not observed. Scale bars, 50 μm. i Cytoplasmic and nuclear fractionation of endogenous NGN3 in P-KO EPs. P-KO EPs accumulated NGN3 in the cytoplasm. The data are presented as the mean ± SEM. WCL, whole-cell lysate; Cyt, cytoplasm; Nuc, nucleus; n.s., not significant. *p < 0.05 (n = 3). See also Supplementary Figs. 4e, 5.
In general, NGN3 protein levels increased in the EP stage and then decreased in the EC stage as hESCs progressed through differentiation (Supplementary Fig. 3a). Next, we investigated whether P-KO is associated with NGN3 stability in the transition from EPs to ECs. Normally, NGN3 protein expression was highest in EPD4 and decreased in ECD1. However, P-KO ECD1 showed greater protein expression than EPD4 (Fig. 3d). As a transcription factor, NGN3 activates the expression of pancreatic developmental genes, including NEUROD1 and PAX68,32,33. We were surprised to find that P-iKO cells treated with dox showed lower expression of NEUROD1 and PAX6 than untreated cells, although we did not observe any differences in the expression levels of the pancreatic developmental genes PDX1, SOX9, and NGN3 (Fig. 3e). In addition, reduced NEUROD1 expression was confirmed at the protein level in dox(+) P-iKO EPs, but the expression of NKX2.2 and NGN3 nontarget genes, such as PDX1, was not affected (Supplementary Fig. 3d–g). To clarify the reduced NGN3 activity in NGN3-positive EPs, NGN3-T2A-EGFP hESCs were generated from P-iKO hESCs (Supplementary Fig. 4). EGFP-reporter revealed that FACS-sorted EGFP-positive EPs showed significantly lower NEUROD1 mRNA expression in the dox(+) condition than in the dox(−) condition. Considering the increased protein levels of NGN3 coupled with the reduced transcript levels of NGN3 target genes (Fig. 3d), it seems that NGN3 does not effectively activate its target genes in P-KO EPs. We expect that either P-KO itself or a lack of arginine methylation suppresses the functionality of NGN3 in hESCs during pancreatic EC development.
Next, we investigated the developmental competence of P-KO EPs to become pancreatic ECs. We found some EC-associated genes (i.e., NKX2.2 and CHGA) with reduced expression in P-KO ECs and others (i.e., PDX1, HNF1b, and MAFA) with normal expression (Fig. 3f). The transcript levels of genes downstream of NGN3 (i.e., NEUROD1, PAX6, and PAX4) were reduced in P-KO ECs (Fig. 3f and Supplementary Fig. 3h, i). Among the pancreatic endocrine hormone genes, we found specific reductions in insulin (INS) and somatostatin (SST) expression at the mRNA and protein levels in P-KO ECs (Fig. 3g, h). Pancreatic polypeptide (PPY) mRNA and protein expression levels were slightly but nonsignificantly increased in P-KO ECs (Fig. 3g, h). Our results imply that PRMT1 KO leads to reduced transcriptional activity of NGN3, thereby impairing the competence of EPs to become β or δ cells in vitro. To determine whether P-KO influences the subcellular localization of NGN3, we examined NGN3 in the cytoplasmic and nuclear compartments of P-iKO EPs. Compared to untreated cells, dox-treated P-iKO EPs showed higher NGN3 protein in whole-cell lysate (WCL) and in the cytoplasm (Cyt) but no difference in the nuclear (Nuc) compartment (Fig. 3i and Supplementary Figs. 4–5). Interestingly, slow migrating bands, presumably phosphorylated forms of NGN3, were observed in the Nuc fraction but not in the Cyt fraction (Fig. 3i). These data suggest that PRMT1-KO causes cytoplasmic NGN3 accumulation and leads to impaired fate choice of EPs.
PRMT1 specifically methylates NGN3 arginine 65
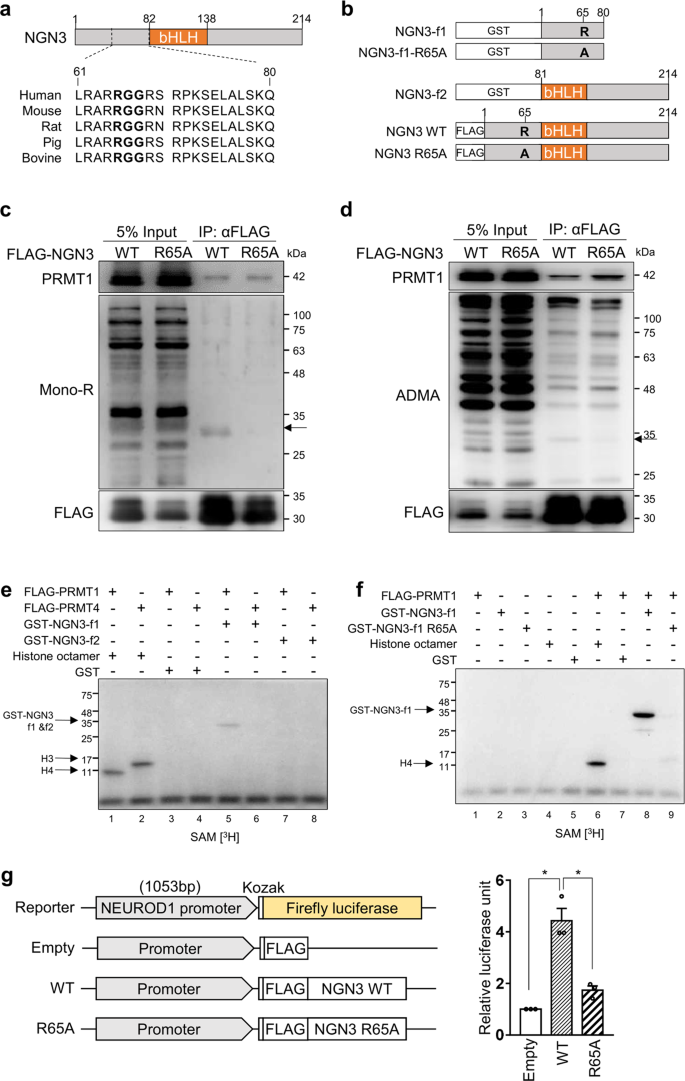
PRMT1 preferentially methylates the arginine residues of RGG/RXR sequences in glycine- and arginine-rich (GAR) motifs in histone and nonhistone proteins15,19,34. According to UniProt (www.uniprot.org), NGN3 has a highly conserved RGG motif (amino acids 65–67) near its basic-helix-loop-helix (bHLH) domain (aa 83–135) (Fig. 4a). To determine whether arginine methylation of NGN3 is regulated by PRMT1, we first generated various FLAG-NGN3 expression vectors. These included vectors expressing intact NGN3, NGN3 fragment 1 (amino acids 1–80, f1), and NGN3 fragment 2 (amino acids 81–214, f2) (Fig. 4b). We also generated NGN3 R65A mutants by replacing arginine 65 (CGC) with alanine (GCC), expecting that PRMT1 would not be able to methylate this site. We then transfected FLAG-NGN3 WT or FLAG-NGN3 R65A mutant plasmids into HEK cells and performed immunoprecipitation followed by western blot analyses with methylated arginine-specific antibodies. We were able to detect mono- and dimethylated arginine bands at the position that corresponds to FLAG-NGN3 protein (arrows) only in the FLAG-NGN3 WT eluate but not in the FLAG-NGN3 R65A eluate (Fig. 4c, d). It is also worth noting that FLAG-NGN3 coimmunoprecipitated endogenous PRMT1 in an arginine 65-independent manner, implying a physical interaction between NGN3 and PRMT1. These results indicate that PRMT1 interacts with and methylates the arginine 65 residue of NGN3.
a An RGG motif (bold) in human NGN3 is conserved across species. bHLH, basic helix-loop-helix domain. b Several human NGN3 constructs for arginine methylation analysis. GST-tagged NGN3 fragments 1 (f1) and 2 (f2) were inserted into pGEX4T-1 for methyl transferase assays. FLAG-tagged NGN3 constructs were inserted into pCAG-FLAG-IPuro for NGN3 overexpression. Arginine 65 (R) of NGN3 was changed to alanine (A) to prevent arginine methylation. c Immunoprecipitation of FLAG-NGN3 with MG132 treatment. A mono-R band (arrow) was detected for FLAG-NGN3 WT but not for the FLAG-NGN3 mutant. mono-R, mono-methylated arginine. d Immunoprecipitation of FLAG-NGN3 without MG132 treatment. A di-methylated ADMA band was detected only with FLAG-NGN3 WT. ADMA, asymmetrical dimethylated arginine. e, f In vitro NGN3 methylation assays. NGN3 arginine 65 was methylated in the presence of [3H]SAM by PRMT1 (e, Lane 5). The R65A NGN3 mutant was not methylated in the presence of [3H]SAM by PRMT1 (f, Lane 8). Histone octamers were used to test FLAG-PRMT1 and FLAG-PRMT4 activities. The GST tag served as a negative control. See also Supplementary Fig. 6. g Luciferase assay for NGN3 activity in HEK cells using the NEUROD1 promoter (1053 bp). The firefly luciferase signal for each experiment was normalized to the Renilla luciferase signal. The data are presented as the mean ± SEM. *p < 0.05 (n = 3). See also Supplementary Fig. 7.
For in vitro methyltransferase assays, we purified FLAG-tagged recombinant PRMT family proteins and the GST-tagged NGN3 protein fragments f1 and f2 (Supplementary Fig. 6a). While both PRMT1 and PRMT4 exhibited intrinsic methyltransferase activity toward histone H4 and H3, respectively, only PRMT1, not PRMT4, could methylate NGN3-f1 (Fig. 4e). In addition, among the nine purified PRMT family proteins tested, only PRMT1 exhibited NGN3 methylation activity (Supplementary Fig. 6b), indicating that PRMT1 is the main arginine methyltransferase for NGN3, which produces mono- and dimethylation states (Supplementary Fig. 6c). In addition, PRMT1 was found to be unable to methylate NGN3-f2 (Fig. 4e), suggesting that PRMT1 methylates the N-terminal region of NGN3. In support of this and consistent with the above cellular analysis, purified R65A-NGN3-f1 was not methylated by PRMT1 (Fig. 4f). Taken together, our cellular and biochemical analyses confirm that PRMT1 specifically methylates NGN3 at arginine 65.
Next, we explored whether arginine methylation of NGN3 influences the expression of its downstream genes. After transfecting NGN3 WT and R65A into HEK cells, we found that the R65A mutant triggered less activation of NEUROD1 than the WT (Supplementary Fig. 7). We then confirmed this reduced transcriptional activity of the R65A mutant via luciferase assay. To do so, we cotransfected HEK cells with each expression vector and a firefly luciferase reporter containing the NEUROD1 promoter32,35. We first confirmed that the expression of each FLAG-tagged version of NGN3 was similar (Supplementary Fig. 7b) and then normalized the firefly luciferase signal to the Renilla luciferase signal. As expected, we observed significantly higher activity triggered by the NGN3 WT transfectants than by the R65A transfectants (Fig. 4g, p < 0.05). To determine the effects of the NGN3 R65A mutant on pancreatic development, H1 PEs were transfected with the NGN3 R65A mutant. Although PEs transfected with NGN3 WT and NGN3 R65A showed similar morphology, the NGN3 mutant group had a lower transcript level of the NEUROD1 gene than the NGN3 WT group, not a lower transcript level of PDX1 (a nontarget gene of NGN3) (Supplementary Fig. 8a). When the NGN3 mutant was transfected into P-iKO PEs, the transcriptional activity of NEUROD1 and PAX6 was reduced in the dox(+) P-iKO PEs compared to the dox(−) P-iKO PEs (Supplementary Fig. 8b). These results indicate that the R65A mutation diminishes NGN3 transcriptional activity. Together, our findings suggest that arginine 65 methylation of NGN3 is essential for transcriptional activity.
Arginine 65 methylation of NGN3 is a prerequisite for its degradation
NGN3 is transiently expressed in pancreatic EPs, rapidly disappearing during pancreatic lineage development12. Therefore, we investigated whether NGN3 arginine methylation is associated with degradation. To answer this question, we transfected P-iKO PEs with an NGN3 expression vector and incubated them for 1–2 d (Fig. 5a). Although ectopic NGN3 expression was significantly reduced 2 days after transfection in P-iKO PEs that were not treated with dox, similar cells treated with dox maintained consistent levels of NGN3 (Fig. 5b). This suggests that PRMT1 is associated with NGN3 degradation in hESCs during pancreatic differentiation.
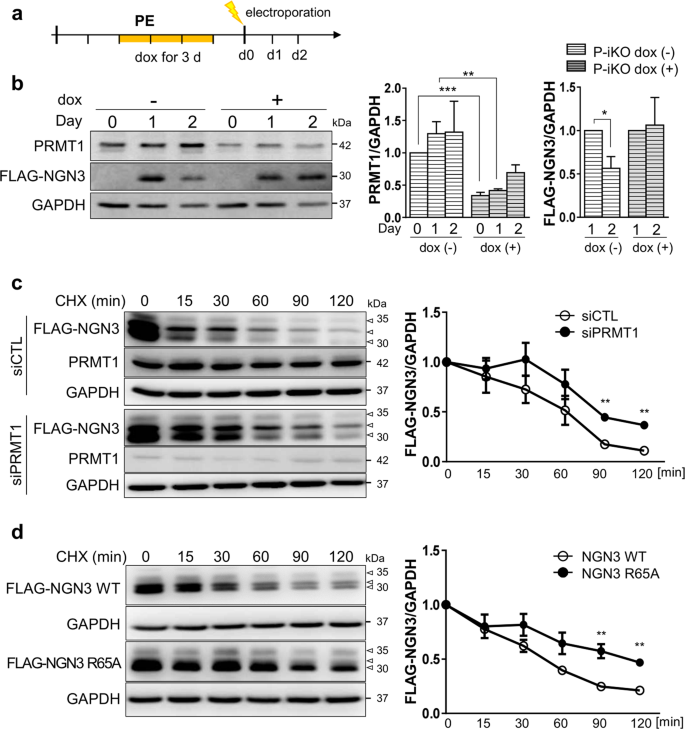
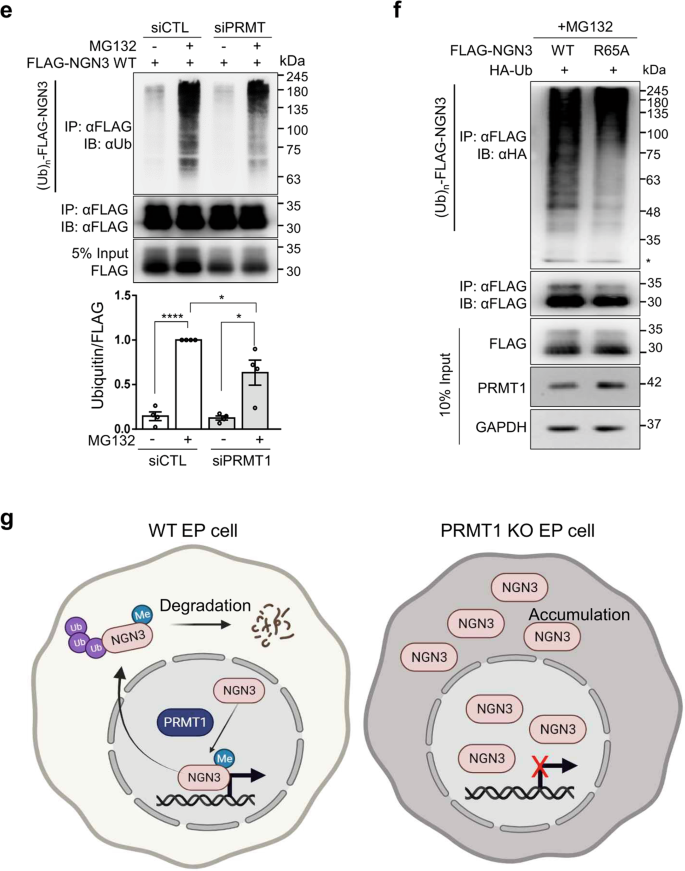
a Protocol for the treatment of P-iKO PEs with dox and electroporation with the FLAG-NGN3 expression vector. b Accumulation of NGN3 protein in P-KO PEs. P-iKO PEs treated with dox(+) showed downregulation of PRMT1 but not FLAG-NGN3. The data are presented as the mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001 (n = 4). c Degradation assay for FLAG-NGN3 in HEK cells treated with PRMT1-siRNA. In the presence of CHX, we observed delayed degradation of FLAG-NGN3 in PRMT1-KD HEK cells. FLAG-NGN3 proteins appear as multiple bands (arrowheads). FLAG-NGN3 levels in cells treated with siPRMT1 compared to those treated with siCTL at the indicated time points after CHX treatment. The data are presented as the mean ± SEM. **p < 0.01 (n = 3). CHX, cycloheximide; siCTL, siRNA control (unspecific sequences). d Delayed degradation of the FLAG-NGN3 R65A mutant in HEK cells. The FLAG-NGN3 R65A mutant was degraded slower than FLAG-NGN3 WT. FLAG-NGN3 R65A mutant levels compared to FLAG-NGN3 WT at the indicated time points after CHX treatment. The data are presented as the mean ± SEM. **p < 0.01 (n = 3). See also Supplementary Fig. 9. e Reduced polyubiquitination of FLAG-NGN3 in PRMT1-KD HEK cells. HEK cells transfected with PRMT1-siRNA showed reduced ubiquitination. The intensity of the (Ub)n-FLAG-NGN3 area was normalized by the αFLAG-NGN3 IP band. Ub, ubiquitin. *p < 0.05, ****p < 0.0001 (n = 4). f Decreased polyubiquitination of the FLAG-NGN3 R65A mutant in HEK cells. g Putative model for the role of arginine 65 methylation of NGN3 in pancreatic EPs. Arginine-methylated NGN3 activates NGN3 target genes such as NEUROD1 in the nucleus and is then rapidly degraded by ubiquitination at the pancreatic EP stage. PRMT1-KO prevents NGN3 arginine methylation, leading to accumulation of NGN3 and reduced activation of its target genes. Ultimately, defective NGN3 arginine 65 methylation prevents the cell fate transition of EPs to ECs. The diagram was generated using BioRender (biorender.com).
To investigate the role of PRMT1 in NGN3 degradation, we transfected HEK cells with PRMT1-specific siRNAs, transfected them with a FLAG-NGN3 expression vector, and then treated them with cycloheximide (CHX) to block protein synthesis. We observed a significant delay in FLAG-NGN3 degradation in the siPRMT1-treated cells compared to the siCTL-treated cells (Fig. 5c, p < 0.01). We also found that FLAG-NGN3 R65A showed a longer half-life than WT (Fig. 5d, p < 0.01). Unlike R65A-NGN3, WT-NGN3 appeared in multiple distinct bands on the western blot (Fig. 5d). In addition, we observed differences in the phosphorylation bands between the FLAG-NGN3 WT and R65A (Supplementary Fig. 9a, arrowheads) that were sensitive to phosphatase treatment (Supplementary Fig. 9b, arrowheads). These results indicate that arginine methylation contributes to NGN3 stability by altering NGN3 phosphorylation. However, phospho-serine-specific antibodies for NGN3 are not yet commercially available, and the exact phospho-serine site influenced by arginine 65 methylation remains elusive.
To examine the correlation between arginine methylation and polyubiquitination of NGN3, we transfected HEK cells with FLAG-NGN3 and siPRMT1 and then treated them with the proteasome inhibitor MG132 (20 μM) for 2 h. In the presence of MG132, the ubiquitinated NGN3 protein significantly accumulated in both groups, but its accumulated level was significantly reduced in the siPRMT1 group compared to the siCTL group (Fig. 5e). R65A-NGN3 also showed weaker ubiquitination bands than WT-NGN3 (Fig. 5f). These results indicate that arginine 65 methylation of NGN3 by PRMT1 is critical for NGN3 degradation in the process of EC development.
- SEO Powered Content & PR Distribution. Get Amplified Today.
- PlatoData.Network Vertical Generative Ai. Empower Yourself. Access Here.
- PlatoAiStream. Web3 Intelligence. Knowledge Amplified. Access Here.
- PlatoESG. Automotive / EVs, Carbon, CleanTech, Energy, Environment, Solar, Waste Management. Access Here.
- BlockOffsets. Modernizing Environmental Offset Ownership. Access Here.
- Source: https://www.nature.com/articles/s12276-023-01035-8